News and blogs
333 results found.
Selinexor recommended in draft final guidance for myeloma
The drug, selinexor has been recommended by NICE in final draft guidance for use in people with myeloma if previous treatment has failed.
22nd Apr 2024

Minreet's marathon for mum with myeloma
Minreet and the Skipping Sikh will run the London Marathon for Blood Cancer UK to give hope to their mum who is receiving treatment for the blood cancer, myeloma
18th Apr 2024

CAR-T therapy Kymriah available for a form of childhood and young-adult leukaemia on the NHS
'Life-changing' blood cancer drug, Kymriah is now available routinely on the NHS for childhood and young-adult B-cell acute lymphoblastic leukaemia on the NHS
11th Apr 2024

How a Yorkshire house led to five pioneering research projects
Last year you raised an incredible £1.95m for Blood Cancer UK in the Omaze Millon Pound House Draw. Find out more about the incredible research projects this is helping to support.
11th Apr 2024

Omaze x Blood Cancer UK Research Fund launched
Blood Cancer UK reveals that the money from their latest partnership with Omaze UK is their largest single donation to date. The money has now been invested into research via the Omaze x Blood Cancer UK Research Fund.
19th Mar 2024

Louis Walsh's blood cancer
Walsh has a rare form of blood cancer, which affects the plasma cells in the blood.
15th Mar 2024

Scotland drug regulators rule blood cancer isn't cost effective
Yescarta, the blood cancer drug is not to be made available on the NHS in Scotland even though it is in England. Blood Cancer UK says it highlights the inequality in access to blood cancer drugs across the UK.
12th Mar 2024
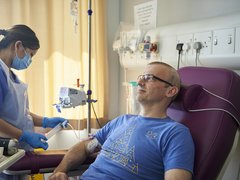
Ambitious £4 million project to develop clinical platform for blood cancer prevention
World Cancer Day sees Blood Cancer UK and The Leukemia & Lymphoma Society announce the start of a transformative five-year, £4 million research project in Cambridge. The work will establish a clinical platform to identify individuals at high risk of certain blood cancers and develop interventions to prevent people from progressing to these diseases.
4th Feb 2024

Non-Hodgkin lymphoma drug, Tepkinly, authorised for use on NHS
Tepkinly (epcoritamab), a treatment for the blood cancer, diffuse large B-cell lymphoma is to be made available on the NHS as a third line treatment.
1st Feb 2024

From despair to hope: The Matthew Wilson Multiple Myeloma Fund
Matthew Wilson establishes the Matthew Wilson Multiple Myeloma Fund (MWMMF) at Blood Cancer UK.
26th Jan 2024

Our response to the Government's plan for UK commercial clinical trials
Refreshing clinical trials activity must benefit people with blood cancer across the UK.
18th Jan 2024

Cards for when "get well soon" doesn't quite cut it
We're thrilled to announced our partnership with artist Vicky Yorke, whose unique card designs and messages draw from her own experience of chronic myeloid leukaemia (CML) – with every purchase supporting life-saving blood cancer research.
12th Jan 2024

Insurance leader Matthew Wilson to be next chair of trustees of Blood Cancer UK
Blood Cancer UK will appoint insurance leader, and advisor to Fairfax Financial and Lloyd’s of London, Matthew Wilson as its next chair of trustees. Matthew, who has been appointed after a robust, independent recruitment process, will formally start his role in early 2024, as current chair John Ormerod’s term comes to an end after 5 years in post.
1st Dec 2023

Amwins Global Risks partners with Blood Cancer UK to raise funds for myeloma
Blood Cancer UK has become Amwins Global Risks’ charity partner, with the partnership expected to raise around £100,000 in the next two years. The partnership will also raise the awareness of blood cancer, which is the third biggest cancer killer in the UK.
20th Nov 2023
Stay updated with our organisational strategy
The latest news on our organisational strategy.
10th Nov 2023

Research links childhood CT scan exposure to increased blood cancer risk
In new research, scientists have linked increased CT scan exposure to an increased risk of blood cancer in younger people.
9th Nov 2023
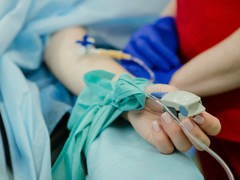
How our research gave vaccine answers to people with blood cancer
As our QResearch project formally comes to a close, we look at how scientists were able to understand the effectiveness of the COVID-19 vaccine for clinically extremely vulnerable people during the pandemic.
2nd Nov 2023

The stolen legacy of Henrietta Lacks
Taken without her consent, Henrietta's 'immortal' cells have had an everlasting contribution to research – while raising important issues around medical ethics and patients' rights. Henrietta's death was the start of a long battle for justice, but 70 years later, Henrietta's family finally received closure.
24th Oct 2023

Non-Hodgkin lymphoma drug authorised for use in the UK
A drug for non-Hodgkin lymphoma, epcoritamab, has been authorised for use in the UK for adults.
24th Oct 2023

This MDS World Awareness Day, four people share their story
October 25 every year is World MDS Awareness Day. This year, four people share their story of living with MDS.
23rd Oct 2023

